Use ALLHiC to assist genome assembly based on HiC data
tags: Bioinformatics Shengxin
Use ALLHiC to assist genome assembly based on HiC data
Genome assembly can be roughly divided into three steps (1) construct contig according to the overlap between sequences, (2) construct contig into scaffold based on the second-generation mate pair library or optical map, (3) sort and adjust scaffold Direction to get the final quasi-chromosome level genome.
The current three-generation sequencing assembly can handle the first and second steps. There are 4 options for upgrading contig/scaffold to the quasi-chromosomal level. One is based on genetic maps, one is based on BioNano DLS optical maps, the other is based on chromosomal homology of closely related species, and the other is HiC. Among them, HiC technology is the simpler of the three. It does not require a high-quality DNA library or a large population, and the results are more accurate and credible.
The schematic diagram of HiC library construction is as follows, what we need is the distance relationship between the two ends of the final paired-end sequencing.

At present, assembling software using HiC data includes LACHESIS, HiRise, SALSA1, 3D-DNA, etc. These software perform well on animal genomes and simple plant genomes, but are not suitable for direct use in polyploid and highly heterozygous species On the assembly. The main reason is the similarity of allele sequences, which makes the contigs of different sets of chromosomes appear false signals, and finally connects the contigs of different sets of chromosomes by mistake. The ALLHiC process recently published in Nature Plants is used to solve the HiC assembly problem of polyploid species and highly heterozygous genomes.
The ALLHiC process at a glance
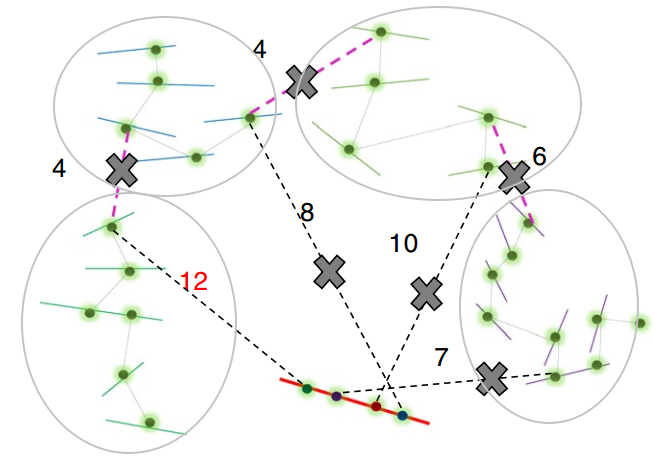
ALLHiC is divided into five steps (see the figure below, Zhang et al., 2019), pruning, partition, rescue, optimization, building, and the required input files are BAM after HiC data comparison and an Allele.ctg.table.

The pruning step is a key step that distinguishes ALLHiC from other software. Therefore, I specifically selected them for introduction. The red solid line is the potential collapse area (due to the similar sequence during assembly, there is no split), and the other color solid lines are different haplotypes (I use light gray ellipses to perform distinguish). The pink dotted line refers to the HiC signal between alleles, and the black dotted line is the HiC signal in the collapsed and uncollapsed regions.
In this step, ALLHiC will filter the HiC signals between alleles in the BAM file according to the Allele.ctg.table provided, and screen out the HiC signals in the collapsed and uncollapsed regions. These signals will be used in the Rescue step to assign unanchored contigs to the grouped contigs.
Software Installation
The installation of ALLHiC is very simple. According to habit, I install the software in~/opt/biosoftunder
mkdir -p ~/opt/biosoft && cd ~/opt/biosoft
git clone https://github.com/tangerzhang/ALLHiC
cd ALLHiC
mv allhic.v0.9.8 bin/allhic
chmod x bin/*
chmod x scripts/*
# Add to environment variables
PATH=$HOME/opt/biosoft/ALLHiC/scripts/:$HOME/opt/biosoft/ALLHiC/bin/:$PATH
export PATHIn addition, ALLHiC also relies on samtools (v1.9), bedtools and matplotlib (v2.0) of the Python 3 environment, which can be done in one step through conda.
conda create -y -n allhic python=3.7 samtools bedtools matplotlibThen check whether the installation is successful
$ conda activate allhic
$ allhic -v
$ ALLHiC_pruneYou may encounter the following error
ALLHiC_prune: /lib64/libstdc .so.6: version `GLIBCXX_3.4.21' not found (required by ALLHiC_prune)This is caused by the GLIBC is too low, but do not try to upgrade GLIBC (you can not afford the consequences), conda provides a relatively new dynamic library, so you can solve the problem by the following methods
export LD_LIBRARY_PATH=$LD_LIBRARY_PATH:~/opt/miniconda3/libALLHiC analysis actual combat
Thanks to the test data provided by Zhang Xingtan, I converted the BAM into the original FastQ format to explain from the beginning. The data is on the Baidu network disk, and you canmy blog Find the link at the corresponding location.
The input file needs to have 4, contig sequence, allelic contig information and two paired-end sequencing data
$ ls -1
Allele.ctg.table
draft.asm.fasta
reads_R1.fastq.gz
reads_R2.fastq.gzStep 1: create an index
samtools faidx draft.asm.fasta
bwa index -a bwtsw draft.asm.fasta The second step: sequence post. The speed limit step for this step isbwa sampe, Because it has no multithreading parameters. If the amount of data is large, you can split the original fastq data first, compare them separately and execute them separatelybwa sampe, And finally merged into a single file.
bwa aln -t 24 draft.asm.fasta reads_R1.fastq.gz > reads_R1.sai
bwa aln -t 24 draft.asm.fasta reads_R2.fastq.gz > reads_R2.sai
bwa sampe draft.asm.fasta reads_R1.sai reads_R2.sai reads_R1.fastq.gz reads_R2.fastq.gz > sample.bwa_aln.sam The third step: SAM preprocessing, remove redundant and low-quality signals, improve processing efficiency
PreprocessSAMs.pl sample.bwa_aln.sam draft.asm.fasta MBOI
# If there is a BAM file
# PreprocessSAMs.pl sample.bwa_aln.bam draft.asm.fasta MBOI
filterBAM_forHiC.pl sample.bwa_aln.REduced.paired_only.bam sample.clean.sam
samtools view -bt draft.asm.fasta.fai sample.clean.sam > sample.clean.bamamong themfilterBAM_forHiC.plThe filter criteria is that the comparison quality is higher than 30 (MQ), only the only comparison (XT:A:U) is retained, the edit distance (NM) is lower than 5, the wrong match is lower than (XM) 4, and there can be no more than 2 Gap (XO, XG)
Step 4 (optional): For polyploid or highly heterozygous genomes, because the sequence similarity of alleles is high, it is very likely that there will be false signals between different sets of genomes, so it is necessaryBuild Allele.ctg.table, Used to filter this kind of false signal.
ALLHiC_prune -i Allele.ctg.table -b sample.clean.bam -r draft.asm.fasta This step generatesprunning.bamFor subsequent analysis
Step 5: This step is to group different contigs according to HiC signals, and the number of groups is determined by-kcontrol. If you skip the fourth step, you can use the result of the third step directlysample.clean.bam
ALLHiC_partition -b prunning.bam -r draft.asm.fasta -e AAGCTT -k 16 This step will generate a series ofprunningFile at the beginning
- Grouping information: prunning.clusters.txt
- Contig corresponding to each group: prunning.counts_AAGCTT.XXgYY.txt:
- The length and count of each contig: prunning.counts_AAGCTT.txt
Step 6: Assign unanchored contigs to existing groups.
ALLHiC_rescue -b sample.clean.bam -r draft.asm.fasta \
-c prunning.clusters.txt \
-i prunning.counts_AAGCTT.txtThis step is based on the groupYY.txt corresponding to the previous prunning.counts_AAGCTT.XXgYY.txt
Step 7: Optimize the order and direction of contig in each group
# Generate .clm file
allhic extract sample.clean.bam draft.asm.fasta --RE AAGCTT
# Optimization
for i in group*.txt; do
allhic optimize $i sample.clean.clm
doneThis step will generate the corresponding groupYY.tour based on groupYY.txt
Step 8: Convert the tour format to fasta format and generate the corresponding agp.
ALLHiC_build draft.asm.fasta This step generates two files, groups.asm.fasta and groups.agp. Among them, groups.asm.fasta is the result we need.
Step 9: Construct a chromatin interaction matrix and evaluate the results based on the heat map
samtools faidx groups.asm.fasta
cut -f 1,2 groups.asm.fasta.fai > chrn.list
ALLHiC_plot sample.clean.bam groups.agp chrn.list 500k pdf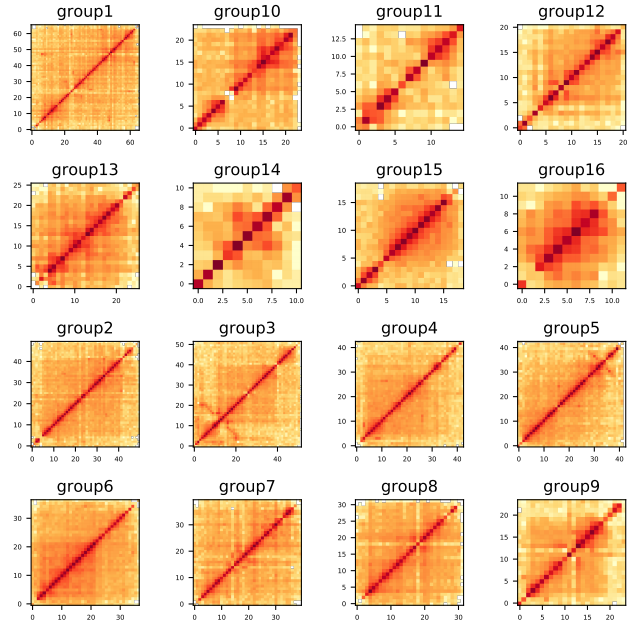
Several precautions for using ALLHiC:
- ALLHiC depends on the initial contig. If the ratio of chimeric sequence to collapsed sequence is too high, the result of ALLHiC will be inaccurate. According to the article, ALLHiC can handle
~10%The mosaic ratio of~20%The ratio of collapse. Therefore, it is best to use assembly software similar to Canu that can distinguish haplotypes. - The sequence similarity between haplotypes should not be too high, otherwise a large number of non-unique alignments will appear, reducing the available HiC signal
- The construction of Allele.ctg.table requires a relatively close high-quality genome
- Don’t use short contigs, because short contigs have fewer signals and can easily be placed in the wrong area
- The K value should be set according to the actual number of genomes. If you find that some groups in the output result are too large, you can appropriately increase the K value
Reference
-
-
- Zhang, X., Zhang, S., Zhao, Q., Ming, R., and Tang, H. (2019). Assembly of allele-aware, chromosomal-scale autopolyploid genomes based on Hi-C data. Nat. Plants 5, 833–845.
- Zhang, J., Zhang, X., Tang, H., Zhang, Q., Hua, X., Ma, X., Zhu, F., Jones, T., Zhu, X., Bowers, J., et al. (2018). Allele-defined genome of the autopolyploid sugarcane Saccharum spontaneum L. Nature Genetics 50, 1565.
----
Copyright Notice: All articles in this blog are used except for special statementsCreative Commons Attribution-Non-Commercial Use-No Derivation 4.0 International License Agreement (CC BY-NC-ND 4.0)Permission is granted.

Intelligent Recommendation
Genome assembly --- Nanopore data assessment
Genome assembly --- Nanopore data evaluation (Naman mustard nanopore) 1. Download software 2. Software use (1)nanoQC (2)NanoPlot 1. Download software Use Conda to create an environment, downloadnanoqc...
Use RaGOO for genome-assisted assembly
RaGOO Raising chromosomes from contig/scaffold level to chromosome level is the ultimate goal of assembly. We usually use genetic maps, optical maps, and HiC technologies to provide information to sor...
Hicanu for HiFi genome assembly | Different sequencing data applications and genome assembly
introduce Canu is specifically assembled by PACBIO or Oxford Nanopore sequences. CANU is divided into three phases: correction, trimming and assembly. The correction phase will increase the accuracy o...
[4] Ragtag-Genomey-based genome assembly based
Introduction to Ragtag RAGTAG can perform error assembly and correction, scaffold assembly and repair, scaffold merger, etc., a total of four steps: Correct, Scaffold, Patch, Merge. After that, you ca...
Abyss: Bloom filter-based genome assembly software
The mainstream NGS genome assembly software first divides the sequence into kmer, and then obtains the assembled sequence based on the de Bruijn Graph graph theory algorithm. When the program is runni...
More Recommendation
HiC-Pro combat #3D genome # epigenetic
First of all, thanks to jimmy for a very detailed tutorial.HiC data analysis in real-time HiC-Pro This article is the second part of the three-dimensional genome study notes, mainly recording the prob...
Genome Assembly
Genome assembly refers to the use of sequencing methods to generate sequence fragments (ie read) from the genome of the species to be tested, and to splice the fragments according to the overlapping a...
HiC-Pro: HiC efficient data preprocessing tools
This article can behttp://xuzhougeng.top/archives/HiC-Pro-An-optimized-and-flexible-pipeline-for-Hi-C-data-processingFree reading HiC-Pro is a highly efficient tool HiC data preprocessing, can be used...
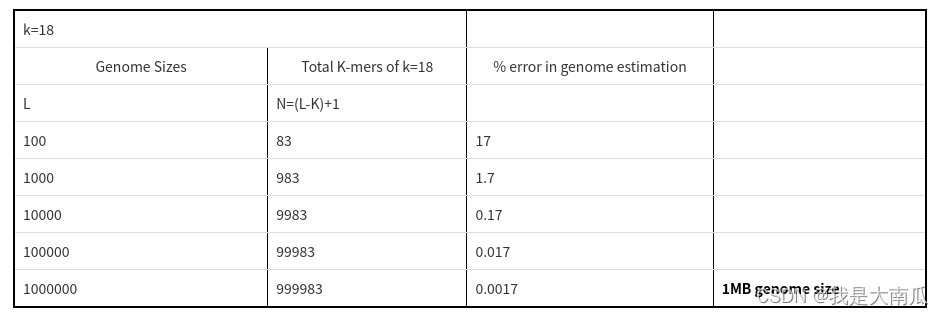
Genome assembly --- Genome Survey (Genome Survey)
Genome assembly --- Genome Survey (Genome Survey) 1. Estimate the principle of genome size 2. Jellyfish software 3. Genomescope Calculate Hybridization 4. GCE software 5. Estimate the size of the Arab...
Using 3D-DNA process to improve genome assembly based on Hi-C
This article can be found inhttp://xuzhougeng.top/Read the original text for free After using the second-generation data or the third-generation data to get the contig, the next step is to raise the c...