Second generation sequencing principle (Illumina)
Fundamental
Fluorescently labeled dNTPs based on reversible termination
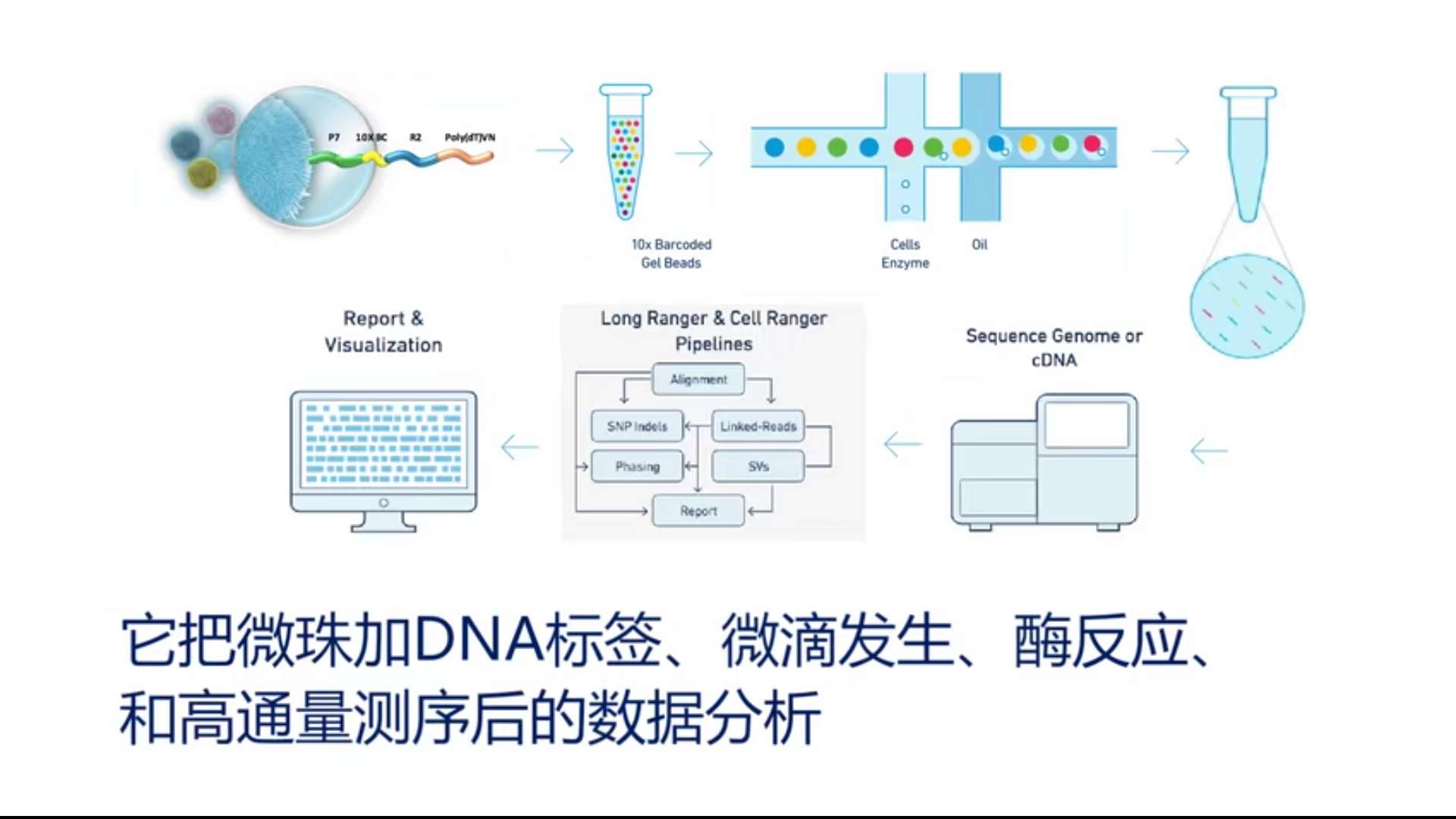
Sample preparationSample Prep
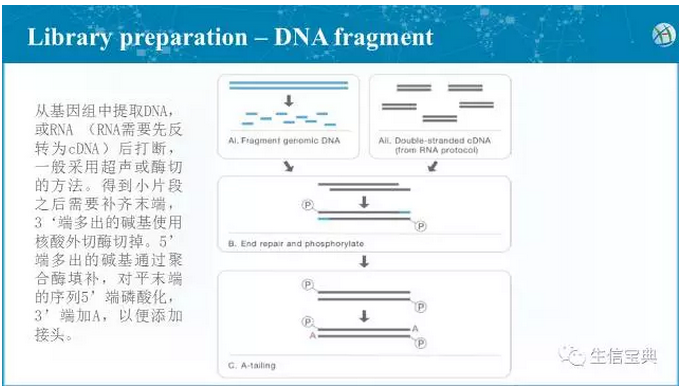
For samples obtained by different experimental methods, the DNA in the sample genome needs to be extracted first, and it is randomly interrupted by ultrasonic waves.
The enzyme was then used to fill the ends, using the Klenow enzyme to add an A base at the 3' end (for ligation of the linker sequence).
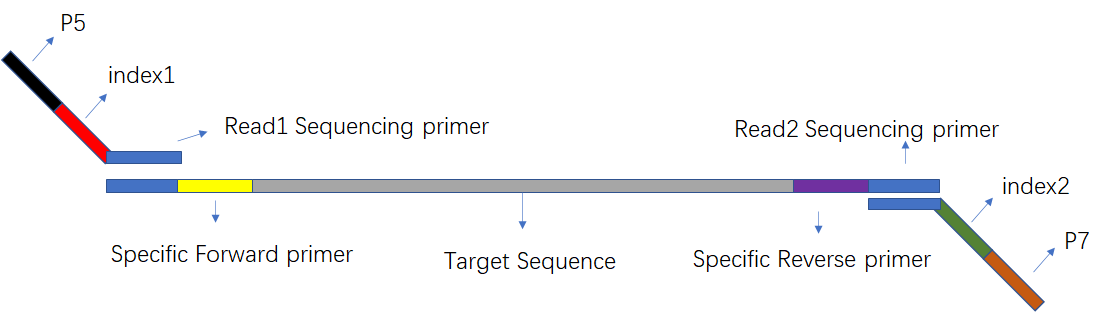
For subsequent amplification, sequencing analysis, specific linker sequences (known for sequencing identification) need to be added to these DNA fragments, and there are roughly three types:
- Sequencing binding site (green)
- Index (red, yellow)
- Flow cell primer complementary sequence (blue, purple)
The collection of DNA fragments after the addition of the linker sequence is called a DNA library, thus completing the sample preparation work.
ClusterCluster Generation
Clustering is the process by which DNA fragments are amplified, which is done in a flow cell. It is a glass carrier with 8 lanes with two DNA primers attached to the inner surface of each channel.

First, the primer will complement the linker sequence of the DNA fragment in the sample and be fixed on the channel surface.
The complementary fragment of the hybrid fragment is generated by the polymerase, and then the NaOH base solution is added, the double-stranded molecule is denatured, and the original template strand (the left strand) is washed away by the liquid in the flow cell.
A neutral liquid is added to neutralize the alkaline solution, and the linker at the other end of the remaining single-stranded copy strand binds to the primer on the surface of the channel to form a single-stranded bridge.
Similarly, with the participation of the polymerase, a complementary strand is generated, eventually forming a double-stranded bridge.
By denaturation, DNA molecules are linearized into two single-stranded copies
They are combined with their own paired primers.
Repeat this cycle while forming millions of clusters. In the process, all DNA fragments are cloned and amplified.
After bridge amplification, the reverse strand is cleaved off, leaving only the forward strand. To prevent the formation of a single-stranded bridge by specific binding, the 3' end is blocked
SequencingSequencing
First, fluorescently labeled dNTPs and enzymes are added to the Flowcell, and the daughter strands are synthesized starting from the primers.
However, the presence of a 3'-end azide group in dNTPs hinders the extension of the daughter strand, which allows only one base to be measured per cycle.
After one base has been synthesized, the Flowcell is flushed with liquid to wash away excess dNTPs and enzymes, and the laser is scanned using a microscope to scan the characteristic fluorescent signal.
The fluorescence emission wavelength together with the signal intensity determines the readout of the base, and one base of all DNA fragments is read simultaneously. In the process of massive parallelism, the image read by the machine is similar to the following
A chemical reagent is added to excise the azide group and the fluorophore, and then the Flowcell is re-introduced with fluorescently labeled dNTPs and enzymes, and a base is synthesized starting from the primer. Repeat this process continuously to complete the first reading.
Since the throughput of the sequencer per sequencing is relatively large, there may be more than one sample per sequence.
To distinguish between each sample and the positive and negative strands, when scientists construct a DNA library, different indices (or barcodes) are added to the linker sequence to distinguish the source.
First, after the first reading is completed, the copied chain will be washed away.
The index fragment primer is introduced and hybridized to the template, and the sequence is read and washed away. The sequence thus read can be aligned with the known index at the beginning to label the measured sequence for subsequent analysis.
Paired-end sequencing has become the mainstream nowadays, which increases the length of sequencing and provides a new method for structural variation analysis. To complete the double-end sequencing, the template chain 3' is first deprotected, the template is folded, and the index fragment is introduced.
Forming a double-stranded bridge with the participation of polymerase
It is then denatured and restored to a single strand. Note that this time the forward chain is cut and washed away, leaving only the reverse chain.

The reverse strand starts with a sequencing primer, similar to a forward strand, and is read after multiple cycles.
data analysis Data Analysis
When sequencing is complete, millions of reads are generated, based on the index constructed at the time of sample preparation. Bases with similar extensions were brought together for each sample. The forward and reverse read pairs generate a continuous sequence. These sequences are constructed by matching the reference genome to achieve complete sequence construction.
Intelligent Recommendation
sc-RNA seq and Illumina sequencing
Course notes 10X genomics 10X genomics technology: Publications in the 10X official website. Quantitative analysis of RNA expression of individual cells. Molecular biology analysis system based on the...
Basic knowledge of second-generation sequencing
Basic knowledge of second-generation sequencing Basic concepts of second-generation sequencing (This is something that every department related to second-generation sequencing must master) FQ data for...
Shengxin Xiaobai's Learning Diary Day 2-NGS Basics Illumina High-throughput Sequencing Principle
May 26, 2019, Sunday, light rain Note: Read the Shengxin Baodian and read the summary of the article. For the original text, please follow the official accountShengxin Baodian, The blog posts referenc...
Second-generation sequencing assembly PK third-generation sequencing assembly
Second-generation sequencing assembly PK third-generation sequencing assembly 2016-07-29 Editor: Nuohe Zhiyuan Three generations of Pacbio sequencing technology With its long read length, no amplifica...
Illumina double-end sequencing (pair end)
This article is from sixu_9days's CSDN blog: The core of illumina sequencing is the sequencing of Sequencing-By-Synthesis using reversible-terminated, fluorescently-labeled dNTPs.SBS) Flowcell is a gl...
More Recommendation
Introduction to some amplicon sequencing projects of Illumina platform
Amplicon sequencing is actually PCR product sequencing, but PCR technology has many variants, so it is collectively called amplicon sequencing. In addition, in layman's terms, high-throughput sequenci...
How to judge the source platform of second generation sequencing
When uploading data, I found an interesting question. I don’t know which sequencing platform the data came from, so I sorted out how to analyze the source of the sequencing data according to the...
How to determine the source of the second generation sequencing platform
Found an interesting problem when uploading data, the data from which they do not know what illumina sequencing platform in, so finishing at how the identity of the source FASTQ line analysis of seque...
SPADES mixed assembly second generation, three generation sequencing data
Guide Spades is published in 2012, a second-generation sequencing assembly software proposed in Journal of Computational Biology, is currently 6200+, and the number of quasiors in the Acer Beca...
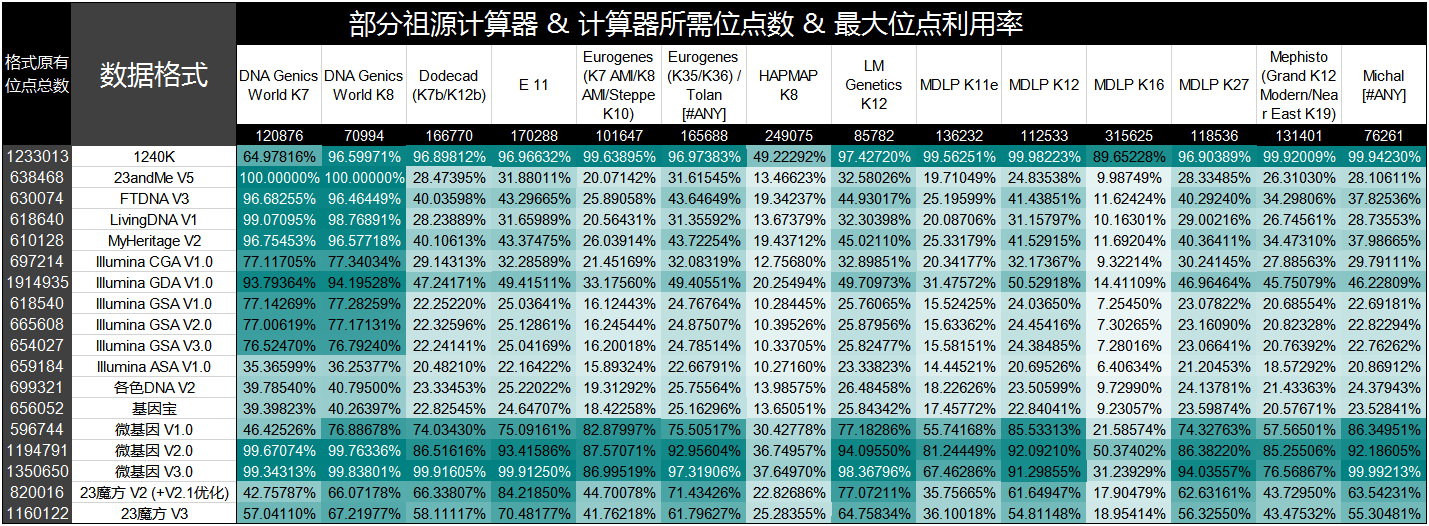
The comparison of several domestic chip sequencing formats and Illumina GSA sites
In recent years, many foreign sequencing companies (such as 23andme, FTDNA, etc.), they will be similar to Illumina Omni Express The chip or other site set is replaced by similar to Illumina GSA Sets ...